How does captivity affect the gut?
Researchers from the Center for Evolutionary Hologenomics have delved into the contentious question of what captivity does to the composition of gut microbiota when compared to wild animals. The results show that the compositional changes are less systematic than previously assumed.
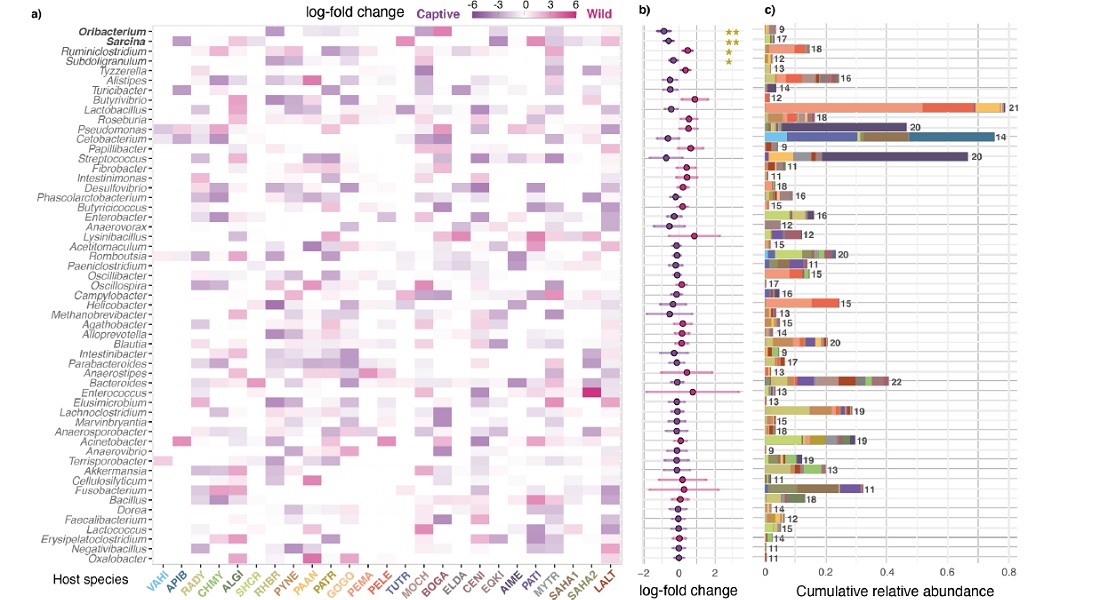
Accepting that the gut microbiota and the host animal are intimately linked, affecting everything from development to well-being, what does the transition from wild to captivity then do to the animal through the consequent changes in the gut microbiota? How does the new environment translate into the composition of the gut? And will the same type of transition - from wild to captivity - reveal a pattern in the compositional changes in and across species?
These are some of the fundamental questions that Associated Professor Antton Alberdi Estibaritz, Assistant Professor Ostaizka Aizpurua Arrieta and Research Assistant Garazi Martin Bideguren from the Center for Evolutionary Hologenomics have addressed for the first time in a new meta-study published in Scientific Reports. In this study they have analysed and compared the gut microbiota profiles of 322 captive to 322 wild animals across 24 different species. The results shake up previous base assumptions and invite further studies.
“So far, there was no systematic meta-analysis on whether and how captivity induces directional changes in the gut microbial communities associated with animals, just a collection of dozens of publications with different study designs and aims. In this study, we have gone through over 200 publications and reanalysed the available data to try to detect overall microbiota variation patterns,” says Antton Alberdi, lead author of the study.
No simple answers
The study shows that the changes in gut microbiota between wild and captive animals is a complex response that does not appear immediately systematic. While the researchers have found that a change in the gut microbiota happens, the changes are diverse and heterogeneous rather than homogenous both in and across species.
“On the one hand, our studies indicate that no rule of thumb can be applied to predict what will happen with the microbiota of animals taken into captivity. It seems clear that, usually, the microbial diversity initially drops, yet after some time, it might fully recover or can even exceed that of wild counterparts. On the other hand, the lack of relevant metadata complicates associating the observed patterns with management features such as the diet, the environment or the use of medicines, which could contribute to increasing our understanding on why microbial communities change in one way or another,” says Antton Alberdi.
Because the responses of gut microbiota compositions are so diverse, there are no simple answers and no broad steps that can be counted on to universally manage captive populations of animals. Instead, as the lack of pattern in responses suggests, managing captive animals should be done from a case-to-case basis, especially looking continuously at whether the changes in gut microbiota has physiological impact on the host animal.
Pitfalls of data assumptions
Interestingly, the researchers have found an increased presence of human-associated microorganisms in captive animals. However, it is impossible to tell if non-human associated microorganisms oust their wild counterparts or simply fill a void that lost wild microbes have left behind. This interaction, the study thereby suggests, should be further explored in a longitudinal study.
Nothing is simple, clearly, when it comes to the gut and the microorganisms inhabiting it. Not only is the presence of human-associated microorganisms difficult to trace precisely, the data in general also has a noticeable overweight of mammals.
“Over 75% of the datasets we have found belong to mammals. There is a huge overrepresentation of mammal-associated microbial communities compared to birds, reptiles and fish, for example. This limits our capacity to decipher whether the observed changes can be linked to general eco-physiological features like energy metabolism (endotherms vs. ectotherms) or environment (land vs. water), and thus generalise conclusions,” Antton Alberdi explains.
Until datasets have been improved, then, the researchers therefore emphasize that data from captive animals should be used with caution when addressing evolutionary questions concerning wild organisms.
Read the full article in Scientific Reports here.
Contact:
Associate Professor Antton Alberdi
